Genomic Services
Source BioScience offers genomic services to Global Pharmaceuticals, Biotechnology organisations, and Life Science and Academic communities.
Using the latest technologies including Illumina Novaseq, NextSeq500, Oxford Nanopore Technologies, Element Biosciences AVITI™ System and MiSeq, Source Genomics is able to provide services for Next Generation Sequencing (NGS), Sanger Sequencing, Bioinformatics, and a range of Molecular Genetic services, operating to ISO9001:2015 standards.
Our team of dedicated highly qualified and experienced scientists will support and update you throughout your project, ensuring you receive the highest quality data and service as well as a quick turnaround.
You will also have a dedicated account manager to support you throughout your project. We offer free consultations, contact us today and one of our account managers will be in touch.
Bigger picture research through multi-omics
Multi-omics is a rapidly expanding field that combines multiple omic technologies to study many layers of biological research simultaneously. Source Genomics is committed to supporting research at every biological level to push forward research and discovery across genomic medicine.
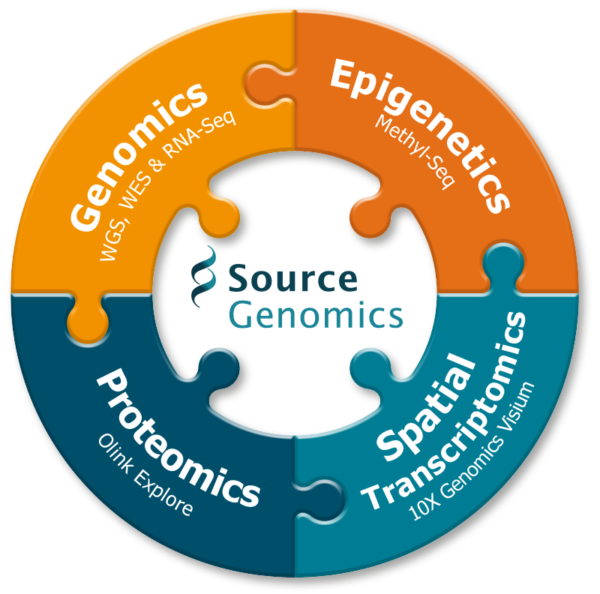
Combining genomic, transcriptomic, proteomic and epigenomic data together, multi-omic research has a wide range of applications to boost research capabilities across all study focus groups:
Biomarker discovery
Identification of novel biomarkers for early disease detection and the development of targeted therapies.
Disease mechanisms
A comprehensive insight into genetic variants, gene expression patterns and protein modifications associated with disease.
Personalised medicine
Understanding individualisation across biological processes associated with disease and advancing tailored treatment approaches in healthcare and pharmaceutical developments.
Drug discovery and development
Identification of potential drug targets and key molecular drivers of disease for an accurate depiction of disease pathways and target intervention systems.
Why Source Genomics?
Benefits of using Source Genomics

Unrivalled
Data Quality

Global provider from
ISO-accredited labs

Rapid Turnaround
Times
Contact us today
For more information or if you would like support with a bespoke project, please contact a member of our team or call
+44 (0) 115 973 9012