Whilst we have streamlined NGS packages for more commonly used applications, we understand that many researchers have specific library preparations and bespoke sample handling requirements and so only require the sequencing of fully prepared NGS libraries.
Whilst we have streamlined NGS packages for more commonly used applications, we understand that many researchers have specific library preparations and bespoke sample handling requirements and so only require the sequencing of fully prepared NGS libraries.
Source Genomics offers run-only sequencing of pre-made libraries on the Illumina NovaSeq, NextSeq and MiSeq instruments, providing a full range of sequencing read modes and data outputs.
Be it 28 Read1 91 Read2 for single cell RNA sequencing all the way through to 2x300 bp for de novo transcriptome assemblies, there is a read mode option for you and with data outputs ranging from 1 million to 10,000 million reads we can tailor to your sequencing project’s needs.
We include data demultiplexing to return sorted sample data with the service and our expert bioinformatics team can provide further data analysis support where required.
A free consultation is provided at the initiation stage of the project and an a dedicated Account Manager will be assigned to provide end-to-end support throughout the project and post data delivery.
Overview
Sequencing package details
As standard we offer entry QC of the provided sequencing pool, PhiX for sequencing QC and increased pool diversity, sequencing on full flow cells using the desired read mode and data output then return of the demultiplexed data via secure FTP server or HDD.
Sequencing instruments: Illumina NovaSeq, NextSeq and MiSeq
Sequencing read modes available: 100, 150, 200, 300, 500 and 600 cycles
- Example: 150bp paired end read mode uses 300 cycle reagents
- Custom read modes available e.g. 28r1 91r2
Sequencing outputs: 1 million to 10,000 million reads per flow cell
The process
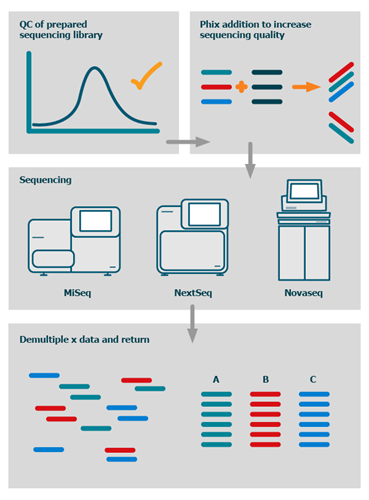
We also offer advanced bioinformatics analysis packages as well as fully bespoke bioinformatics solutions.
For run-only projects, as standard we include the demultiplexing of raw data free of charge providing that library indexing information is provided. However, we can also utilise all relevant bioinformatics pipelines and we offer bespoke solutions if required.
A non-exhaustive list of possible bioinformatics analysis options is below:
- De novo genome assembly
- SNP and InDel analysis and annotation
- Trio/custom comparative analysis
- De novo transcriptome assembly
- Digital or differential gene expressions analysis
- Detection and quantification of gene-fusions and splice variants
- Metagenome taxonomic classification and differential abundance analysis
For a more in-depth discussion on our bioinformatics capabilities please contact your account manager or the bioinformatics team directly.
Genomics@sourcebioscience.com
How to order
Contact us today and one of our skilled account managers will be in touch with a free consultation including further information and pricing details.
Payment options:
Payment can be made by credit card or purchase order number.
FAQ's:
View our frequently asked questions for more information.
Sample requirements:
Please click here to view sample requirements.
How to Order
Contact us today and one of our skilled account managers will be in touch with a free consultation including further information and pricing details.
Next Generation Sequencing
Source BioScience is one of Europe’s leading providers of commercial sequencing, offering Next Generation Sequencing services from our ISO accredited laboratories. We offer NGS services on the most prominent platforms including Illumina’s NovaSeq, NextSeq and MiSeq.