Difference of species composition16s rRNA gene is a highly conserved component which contains abundant taxonomic information. It is the most widely used gene marker for genus and species identification and taxonomic significance in bacteria and archaea. In Metagenomics, the 16S gene amplicons obtained from PCR can be used to deduce taxonomic identifications based upon bioinformatics alignments.
The 18S rRNA is more commonly used in fungi for phylogenetics since it has more hypervariable domains than 16S. However, the ITS (Internal Transcribed Spacer) region (including 5.8S), is more variable and hence more suitable as the genetic marker for measuring intraspecific genetic diversity.
16S/18S/ITS Amplicon Sequencing is a well-established method for microbial identification and phylogeny studies of samples from complicated microbiomes or environments.
Applications
• Species identification
• Gut microbial
• Environment research
• Microbiota diversity
A free consultation is provided at the initiation stage of the project and an a dedicated Account Manager will be assigned to provide end-to-end support throughout the project and post data delivery.
Overview
Sequencing package details
The targeted metagenomics sequencing package utilises a 2-step PCR library preparation approach to amplify your choice of 16S, 18S or ITS regions and incorporate unique dual indexes and Illumina adapters into the library for sequencing. Samples are sequenced using the Illumina MiSeq instrument in the 300bp paired-end read mode to a guaranteed minimum of 40,000 read pairs per sample.
By using a 2-step PCR approach, the package can also be easily adapted for any amplicon with complimentary overhangs to undergo the 2nd index PCR step. Please consult our Genomics team for more information.
Our standard targeted metagenomics package includes:
- Sample entry QC
- Library preparation via 2-step PCR protocol
- Illumina 300bp PE sequencing, guaranteed >40,000 read pairs per sample
- Return of FastQ files via secure FTP or HDD
Optional extras:
- DNA Extraction
- Bioinformatics analysis
Custom Target
Samples for 2nd index PCR only should be prepared using primers with overhangs. Primers should be designed as below and resulting PCR product should form a discrete band on a gel with length of 450-550bp.
Forward primer: 5’ TCGTCGGCAGCGTCAGATGTGTATAAGAGACAG‐[locus-primer]
Reverse primer: 5’ GTCTCGTGGGCTCGGAGATGTGTATAAGAGACAG‐[locus-primer]
Samples will then undergo 2nd index PCR step and subsequent sequencing to 40,000 read pairs.
BioInformatics Taxonomical Classification
Taxonomic classification of mixed samples to provide a profile of microbial diversity
By sequencing conserved regions of ribosomal RNA – 16S for prokaryotic and 18S for eurkaryotic, or the ITS region for fungal samples – taxonomic classification can be achieved. The analysis involves trimming raw sequencing reads, picking operational taxonomic units (OTUs), matching these against a database of choice, such as GreenGenes or Silva, to obtain a break-down for each sample at each of 7 levels: kingdom, phylum, class, order, family, genus and species. You will receive raw sequencing files (fastq), text files detailing OTUs and taxonomic classifications as well as bar charts for each sample. A bioinformatics report will provide further detail about the tools used and a summary of results.
The process
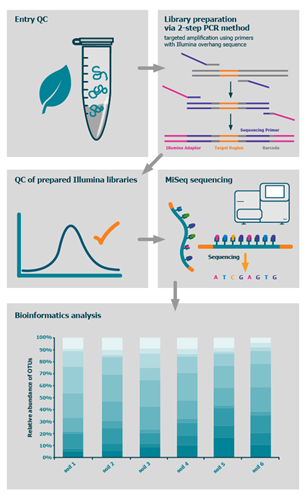
How to order
Contact us today and one of our skilled account managers will be in touch with a free consultation including further information and pricing details.
Payment options:
Payment can be made by credit card or purchase order number.
FAQ's:
View our frequently asked questions for more information.
Sample requirements
Targeted metagenomics package requires 100ng of gDNA (OD260/280 ratio ranging from 1.8-2) at a minimum concentration of 10 ng/μl and in a minimum volume of 10 μl. Click here for more details.
Sequencing Platform
|
Platform Type |
Illumina NovaSeq 6000 |
|
Read Length |
Paired-end 250 bp |
|
Recommended Sequencing Depth |
30K/50K/100K reads |
How to order
Contact us today and one of our skilled account managers will be in touch with a free consultation including further information and pricing details.
Next Generation Sequencing
Source BioScience is one of Europe’s leading providers of commercial sequencing, offering Next Generation Sequencing services from our ISO accredited laboratories. We offer NGS services on the most prominent platforms including Illumina’s NovaSeq, NextSeq and MiSeq.